The most common artifact is powerline noise (50 Hz or 60 Hz) that is usually induced to the acquisition systems. Further, data contaminated with artifactual signals from several types including artificial or physiological sources. delta, alpha & beta bands oscillations) can be identified and characterized. Common iEEG graphoelements of physiological activity (e.g. This approach could also prove useful for supervised adaptive retraining in active learning and expert-in-the-loop scenario based on expert’s review of the data yielding false positive or false negative classifications. We recently demonstrated the ability to temporally localize graphoelements that drive the final classification and make visual review and interpretation of raw EEG recordings possible 8. The inability to dissect the CNN decision process makes it less interesting for understanding fundamental neurophysiology, and ultimately for usage in clinical practice. We have recently demonstrated robust generalization of automated detection algorithms for artifact classification using training and testing datasets collected from different institutions, acquisition systems, under different measurement conditions 7, 8.Īutomated processing and data mining with Convolutional Neural Networks (CNN) are powerful, but the interpretation of particular classifications and correlation with known iEEG waveforms is difficult. The generalizability of an automated artifact detection method will enable broader utilization and extension for any retrospective and prospective iEEG dataset, but this has received little attention. In recent years, a variety of methods using machine learning techniques and deep learning techniques for iEEG processing emerged with impressive results 6, 7, 8. The automatic classification of artifacts and segmentation of iEEG recordings is recognized as a challenging task, and many interesting studies have been published addressing the challenges 4, 5. If reliable automated methods, can be developed they can easily and reliably mine the data and crop the segments of the data with the iEEG features of interests. It is very burdensome for a human operator to manually classify hundreds of channels for data spanning days to weeks.
to focus on the data that are clinically important like seizures or other brain states for research purposes. The acquisition of large datasets has driven the development of improved data preprocessing tools that enable extraction of important application specific data segments i.e. The amount of collected data is rapidly increasing and advancement in data compression, storage, visualization, and automated processing of data is important and has received significant attention 2, 3. In addition to the clinical utility for mapping epileptic brain by localization of seizures and interictal epileptiform transients these data enable a wide range of neuroscience research activities. The improvement of EEG acquisition systems, data storage, and surgical techniques allows for large scale data collection spanning over multiple days to weeks, recording from hundreds of electrodes with sampling rates reaching up to 32 kHz in research settings 1. Currently, the visual inspection and artifact rejection of the data is standard pre-processing procedure that must be done prior to evaluating of epileptic seizure onset zones. The iEEG signals are directly measured from cortical and deep brain structures, e.g. Intracranial electroencephalography (iEEG) is an invasive procedure commonly used for localization of epileptic seizure onset zones in patients with drug resistant epilepsy. Along with the data, we also propose a statistical method that is recommended for comparison of candidate classifier performance utilizing out-of-institution/out-of-patient testing.


The dataset published here should support and facilitate training of generalized machine learning and digital signal processing methods for intracranial EEG and promote research reproducibility.
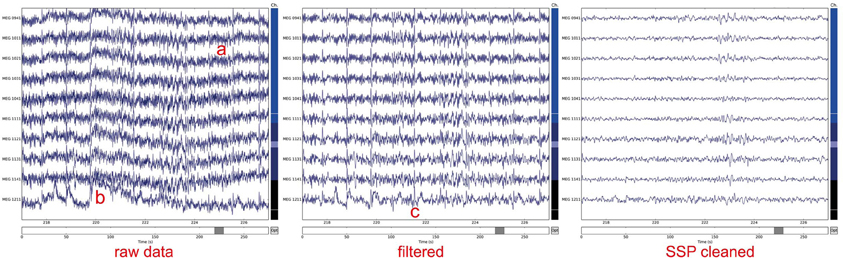
The dataset contains intracranial EEG that are labeled into three groups: physiological activity, pathological/epileptic activity, and artifactual signals. Anne’s University Hospital Brno, Czech Republic. In this study, we publicly share annotated intracranial EEG data clips from two institutions: Mayo Clinic, MN, USA and St. However, the size of EEG data recordings is rapidly increasing with a trend for higher channel counts, greater sampling frequency, and longer recording duration and complete reliance on visual data review is not sustainable. Historically the classification of EEG into physiological, pathological, or artifacts has been performed by expert visual review of the recordings. EEG signal processing is a fundamental method for neurophysiology research and clinical neurology practice.
